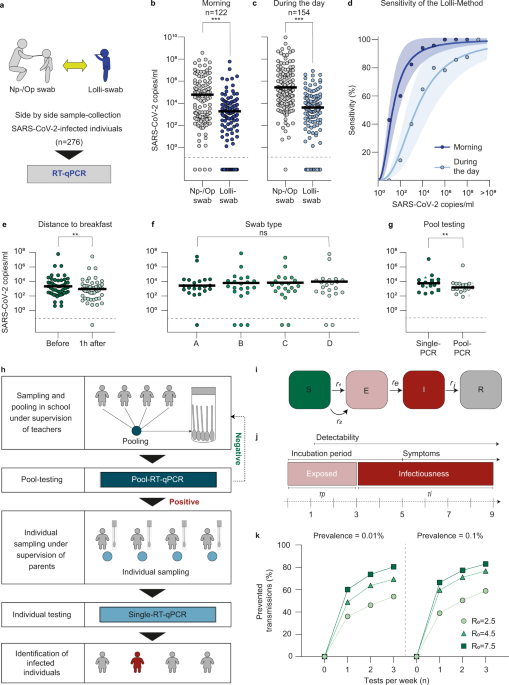
Effective high-throughput RT-qPCR screening for SARS-CoV-2 infections in children | Nature Communications
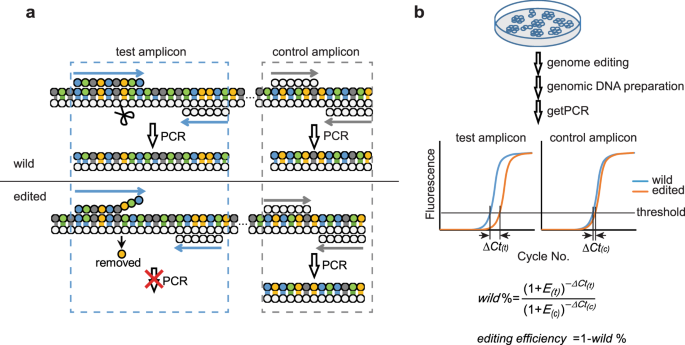
A qPCR method for genome editing efficiency determination and single-cell clone screening in human cells | Scientific Reports
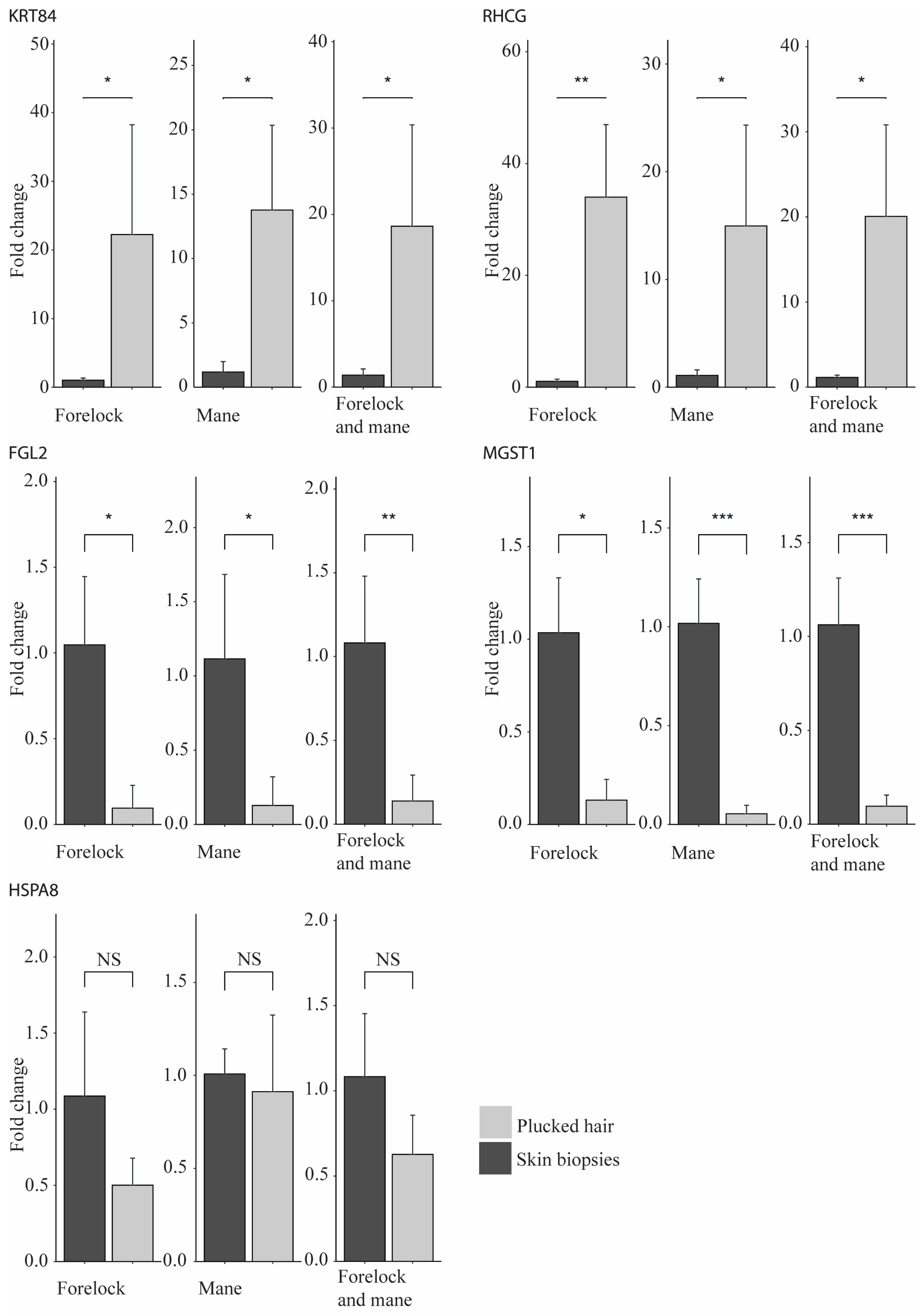
IJMS | Free Full-Text | The Enrichment of Specific Hair Follicle-Associated Cell Populations in Plucked Hairs Offers an Opportunity to Study Gene Expression Underlying Hair Traits
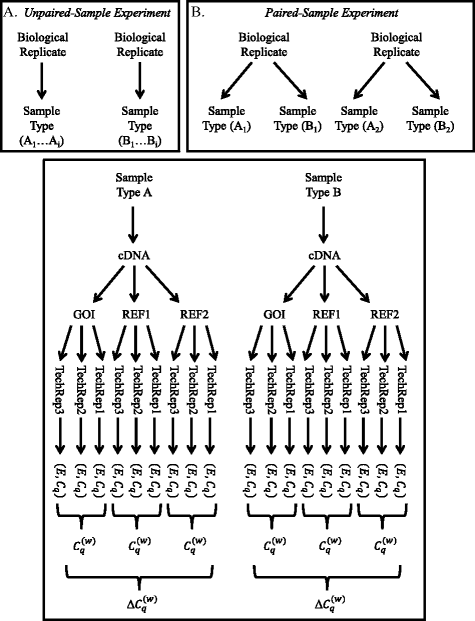
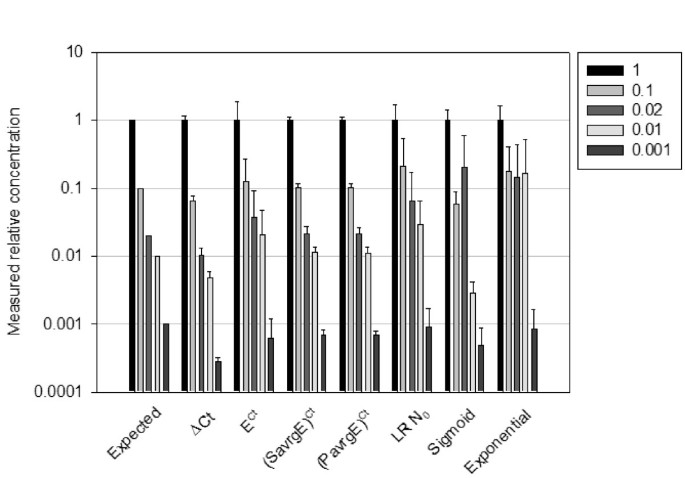
A common base method for analysis of qPCR data and the application of simple blocking in qPCR experiments | BMC Bioinformatics | Full Text
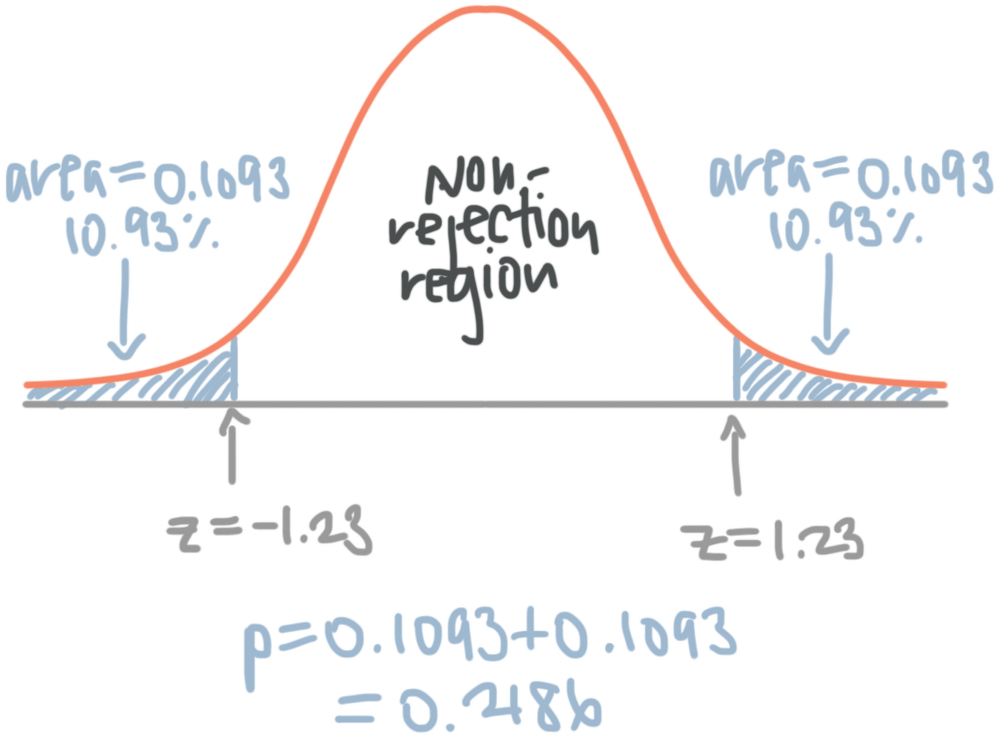
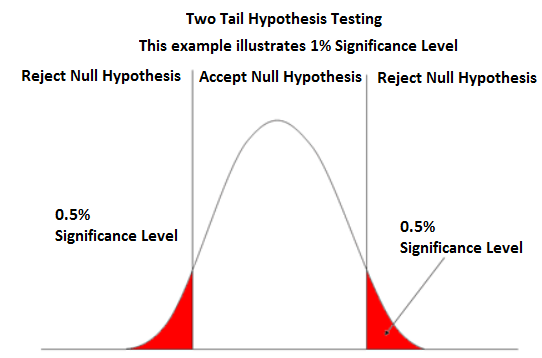
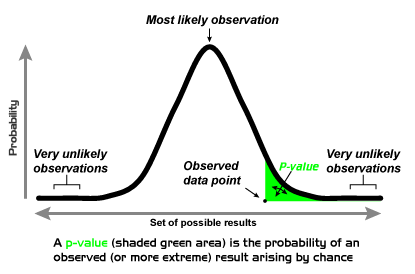
The p-value and rejecting the null (for one- and two-tail tests) — Krista King Math | Online math help
Optimal use of statistical methods to validate reference gene stability in longitudinal studies | PLOS ONE
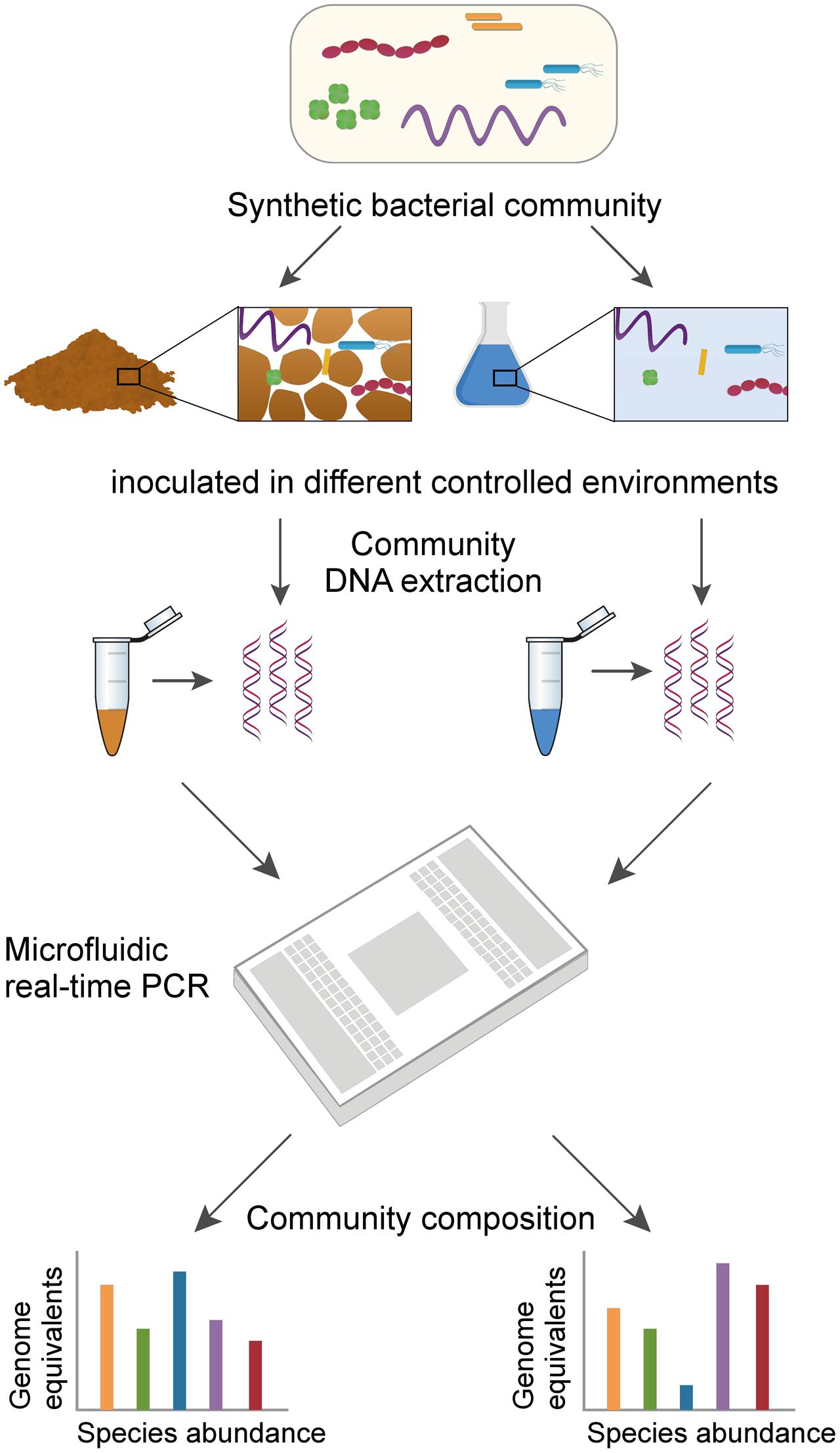
Frontiers | Resolving Species Level Changes in a Representative Soil Bacterial Community Using Microfluidic Quantitative PCR

Clinical performance evaluation of SARS-CoV-2 rapid antigen testing in point of care usage in comparison to RT-qPCR - eBioMedicine

An integrated cell culture reverse transcriptase quantitative PCR (ICC-RTqPCR) method to simultaneously quantify the infectious concentrations of eight environmentally relevant enterovirus serotypes - ScienceDirect